Evolution of cooperation
University of Oxford
https://scholar.google.com/citations?user=e3c1TdffPOcC
Read the preprint here www.biorxiv.org/content/10.1...
Millions of species here we come!
(10/10)
Read the preprint here www.biorxiv.org/content/10.1...
Millions of species here we come!
(10/10)
Just provide the complete set of amino acid sequences for your species
If you prefer a specific tree or alignment tool, it's easy to customise
We also provide rich outputs like gene duplications and comparative genomics stats (9/10)
Just provide the complete set of amino acid sequences for your species
If you prefer a specific tree or alignment tool, it's easy to customise
We also provide rich outputs like gene duplications and comparative genomics stats (9/10)
We now use gene tree–species tree reconciliation to refine orthogroups
This catches cases where distinct orthogroups were mistakenly fused (8/10)

We now use gene tree–species tree reconciliation to refine orthogroups
This catches cases where distinct orthogroups were mistakenly fused (8/10)
We tested using the gold standard Quest for Orthologs benchmarking service
OrthoFinder scored highly across the board (7/10)

We tested using the gold standard Quest for Orthologs benchmarking service
OrthoFinder scored highly across the board (7/10)
We benchmarked orthogroups using the OrthoBench dataset
OrthoFinder came out on top (6/10)
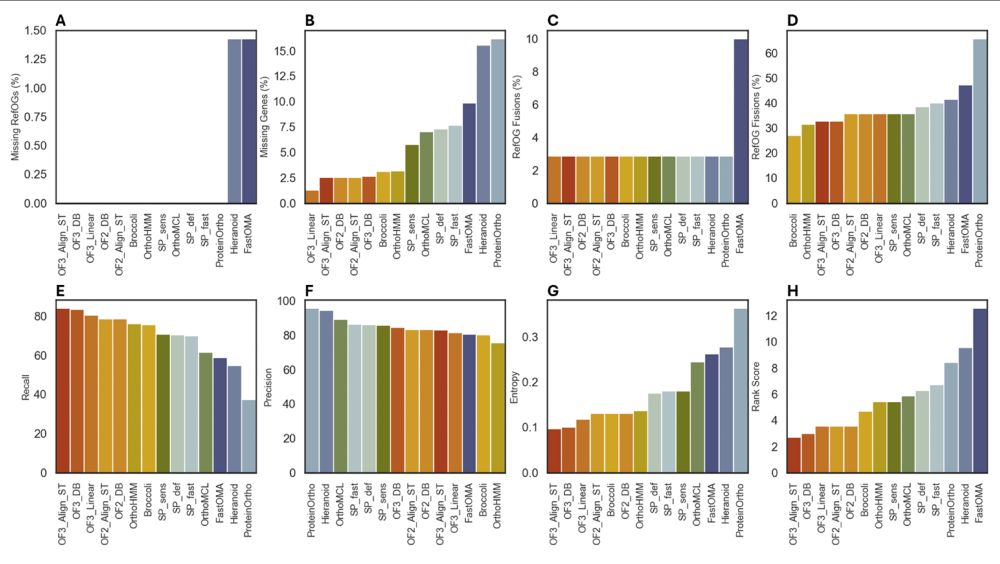
We benchmarked orthogroups using the OrthoBench dataset
OrthoFinder came out on top (6/10)
We benchmarked OrthoFinder against other widely used orthology tools
OrthoFinder is the only method able to analyse >1000 species within our time cutoff (5/10)

We benchmarked OrthoFinder against other widely used orthology tools
OrthoFinder is the only method able to analyse >1000 species within our time cutoff (5/10)
Next, we sample representative sequences from each orthogroup to build profiles
Genes from new species are then matched to these profiles to assign them to orthogroups
We avoid the costly all-vs-all step that kills scalability (4/10)
Next, we sample representative sequences from each orthogroup to build profiles
Genes from new species are then matched to these profiles to assign them to orthogroups
We avoid the costly all-vs-all step that kills scalability (4/10)
This becomes painfully slow as datasets grow
We needed a better way (3/10)
This becomes painfully slow as datasets grow
We needed a better way (3/10)
That’s a huge opportunity, but also a major challenge
How can we ramp up scalability without compromising accuracy?
That’s exactly what we set out to solve in this update (2/10)
That’s a huge opportunity, but also a major challenge
How can we ramp up scalability without compromising accuracy?
That’s exactly what we set out to solve in this update (2/10)
Buff tip, peach blossom, brimstone!
Buff tip, peach blossom, brimstone!


