Excited to share SEAM: Systematic Explanation of Attribtuion-based Mechanisms. SEAM is an explainable AI method that dissects cis-regulatory mechanisms learned by seq2fun genomic deep learning models.
Led by @EESetiz
1/N 🧵👇

Excited to share SEAM: Systematic Explanation of Attribtuion-based Mechanisms. SEAM is an explainable AI method that dissects cis-regulatory mechanisms learned by seq2fun genomic deep learning models.
Led by @EESetiz
1/N 🧵👇

www.biorxiv.org/content/10.1...

www.biorxiv.org/content/10.1...
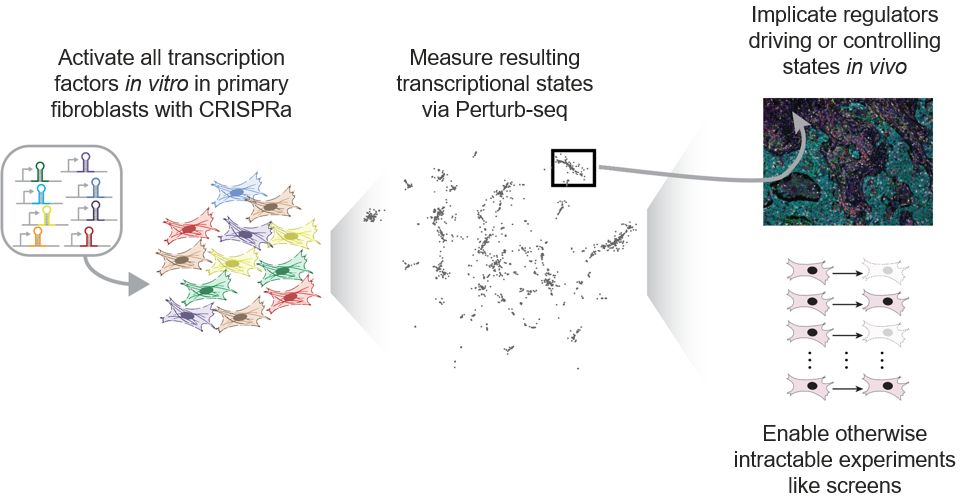
Led by @sherrynyeo.bsky.social, @erinmayc.bsky.social, and friends, we continue our journey to find viral DNA in our favorite place-- the overlooked and discarded reads in existing data! 1/

Led by @sherrynyeo.bsky.social, @erinmayc.bsky.social, and friends, we continue our journey to find viral DNA in our favorite place-- the overlooked and discarded reads in existing data! 1/
genomebiology.biomedcentral.com/articles/10....
genomebiology.biomedcentral.com/articles/10....
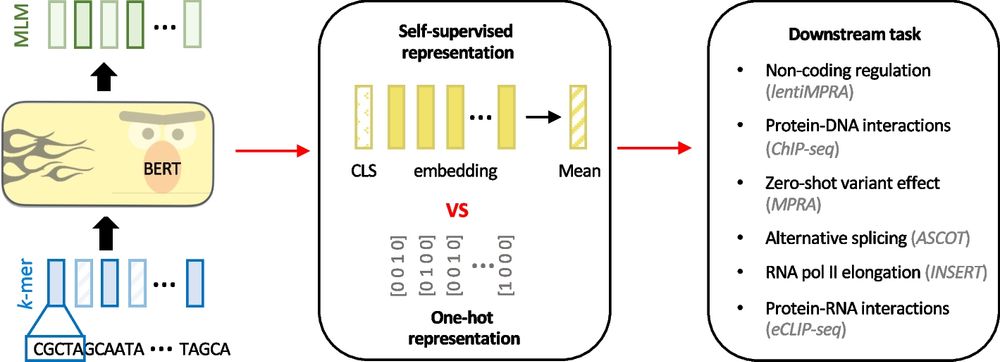
Paper: doi.org/10.1101/2025...
Code: github.com/morrislab/mR...
A 🧵

Paper: doi.org/10.1101/2025...
Code: github.com/morrislab/mR...
A 🧵

www.biorxiv.org/content/10.1...
How does enhancer location within a TAD control transcriptional bursts from a cognate promoter?
Experiments by Jana Tünnermann and modelling by Gregory Roth

www.biorxiv.org/content/10.1...
How does enhancer location within a TAD control transcriptional bursts from a cognate promoter?
Experiments by Jana Tünnermann and modelling by Gregory Roth
shorturl.at/2LHbw
shorturl.at/2LHbw
📄Haplotype analysis reveals pleiotropic disease associations in the HLA region
🧑🤝🧑 @jkpritch.bsky.social @finngen.bsky.social & colleagues

📄Haplotype analysis reveals pleiotropic disease associations in the HLA region
🧑🤝🧑 @jkpritch.bsky.social @finngen.bsky.social & colleagues
Below is a brief description of the major findings. Check the full version of the paper for more details: www.nature.com/articles/s41588-025-02248-5
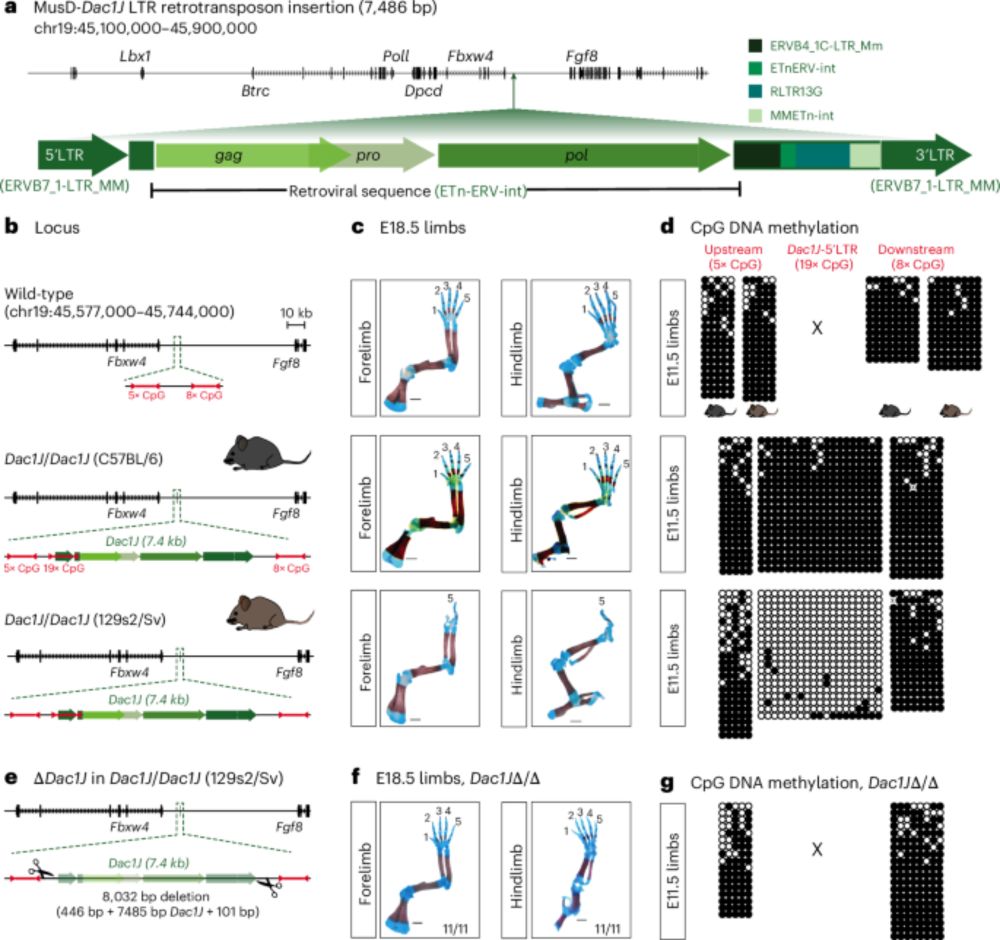
Below is a brief description of the major findings. Check the full version of the paper for more details: www.nature.com/articles/s41588-025-02248-5

medrxiv.org/content/10.1101/2025.06.24.25330216
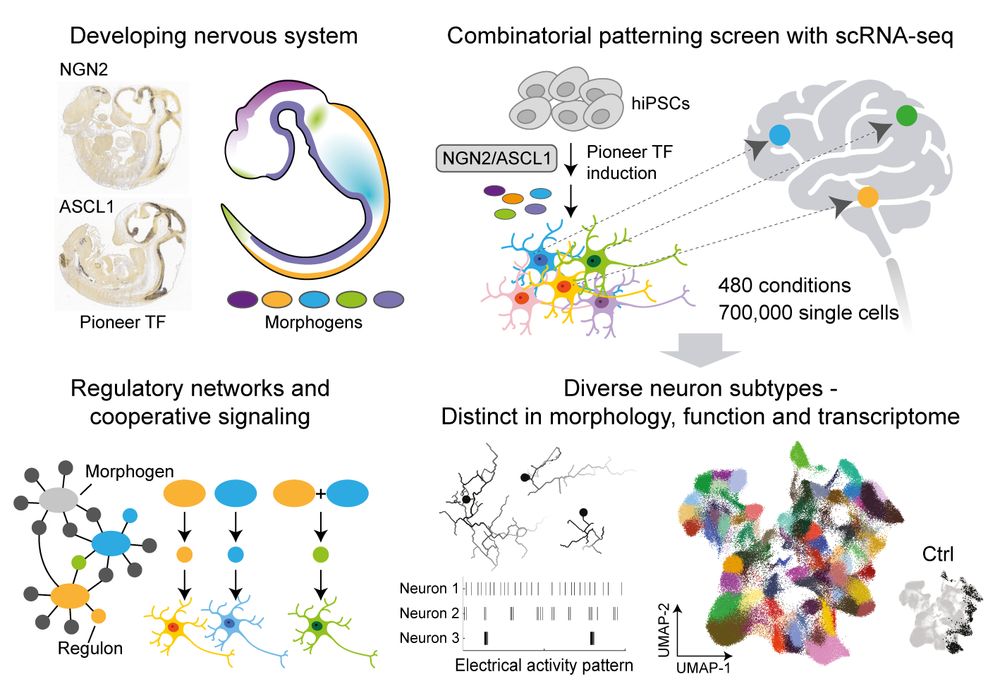
The main output from my PhD is finally public and we’re SUPER excited about the findings! If you’re interested in what we learnt about IBD with a massive 700+ sample sc-eQTL dataset of the gut, read on!

medrxiv.org/content/10.1101/2025.06.24.25330216
The main output from my PhD is finally public and we’re SUPER excited about the findings! If you’re interested in what we learnt about IBD with a massive 700+ sample sc-eQTL dataset of the gut, read on!
More than 2700 human 3′UTRs are highly conserved. These 3′UTRs are essential components in mRNA templates, as their deletion decreases protein activity without changing protein abundance. Highly conserved 3′UTRs help the folding of proteins with long IDRs.
www.biorxiv.org/content/10.1...
More than 2700 human 3′UTRs are highly conserved. These 3′UTRs are essential components in mRNA templates, as their deletion decreases protein activity without changing protein abundance. Highly conserved 3′UTRs help the folding of proteins with long IDRs.
www.biorxiv.org/content/10.1...


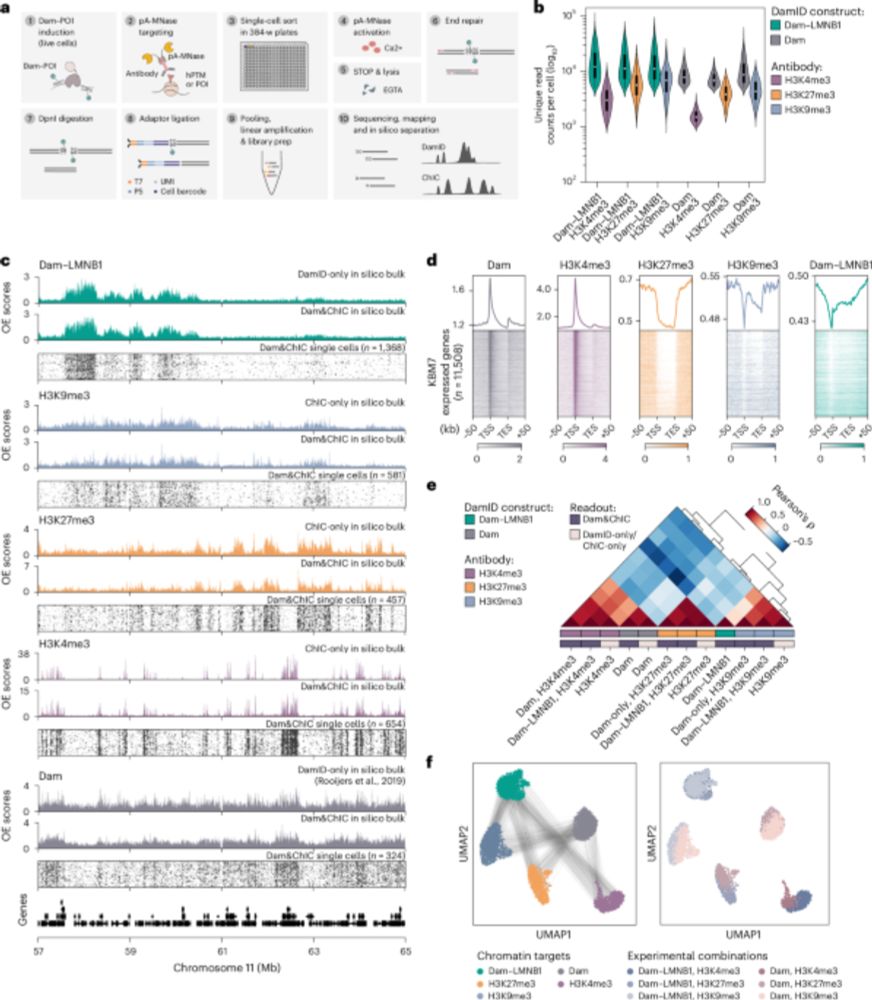
But how do they REALLY work?
New paper with many contributors here @berkeleylab.lbl.gov, @anshulkundaje.bsky.social, @anusri.bsky.social
A 🧵 (1/n)
Free access link: rdcu.be/erD22
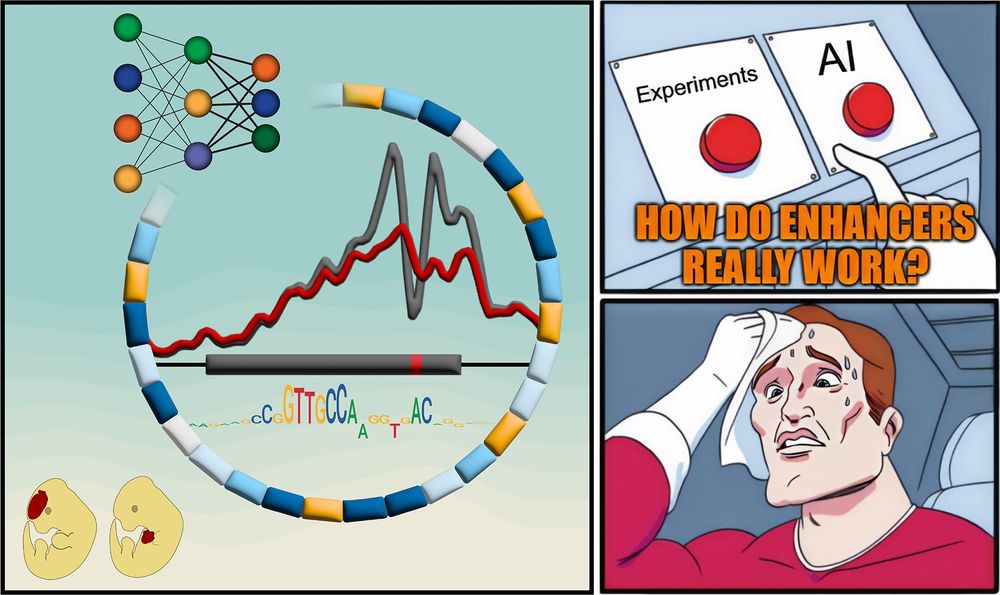
But how do they REALLY work?
New paper with many contributors here @berkeleylab.lbl.gov, @anshulkundaje.bsky.social, @anusri.bsky.social
A 🧵 (1/n)
Free access link: rdcu.be/erD22


