Our team at the Cardiovascular Proteomics group (CNIC) developed PACREDOX, a new method that combines global + cysteine-redox proteomic profiling in a fast, robust way.
👉 Published in Analytical Chemistry:
pubs.acs.org/doi/full/10....Contact us if you want to get the protocol 🗒️
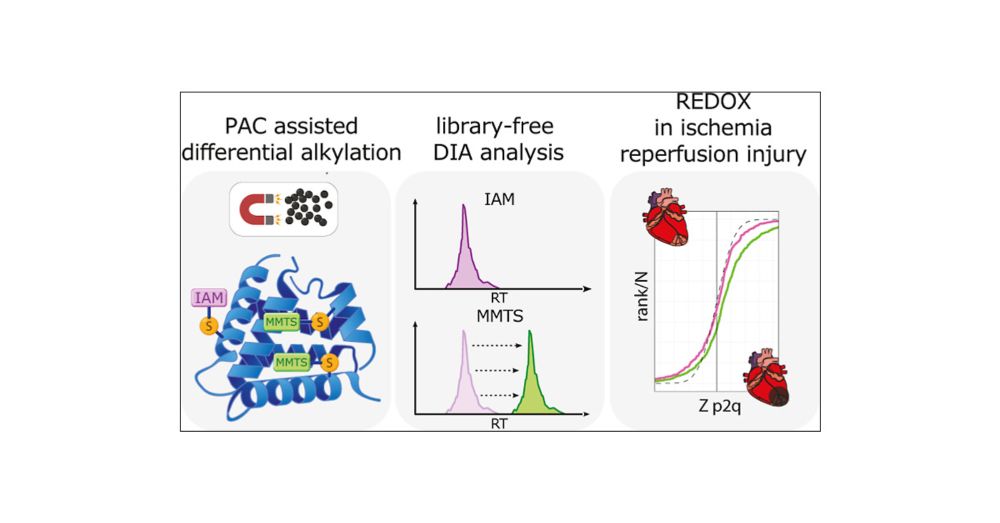
Benchmark for Quantitative Global and Redox Proteomics Analysis by Combining Protein-Aggregation Capture and Data Independent Acquisition
Oxidative damage plays a critical role in various diseases including cardiovascular and neurological disorders. Thiol redox reactions, acting as oxidative stress sensors, influence protein structure and function. Redox proteomics, based on the differential alkylation of cysteine sites followed by mass spectrometry, enables the comprehensive analysis of thiol redox status in cells and tissues. However, these approaches require extensive sample manipulation and are not compatible with data-independent acquisition techniques. Here, we introduce PACREDOX, an innovative strategy based on protein aggregation capture (PAC), and demonstrate its compatibility with library-free DIA. Compared with traditional methods such as FASILOX, PACREDOX reduces preparation time and costs while maintaining thiol and proteome coverage. To enable library-free DIA, we corrected in silico spectral libraries in DIA-NN using experimental retention time data from methylthiolated-Cys peptides. PACREDOX with DIA was benchmarked against FASILOX in a myocardial infarction model, yielding the same biological insights, while enhancing peptide and protein coverage. Our results underscore the potential and efficiency of this methodology for studying oxidative damage. Overall, PACREDOX offers an automatable, high-throughput, and cost-effective strategy for redox proteomics.






