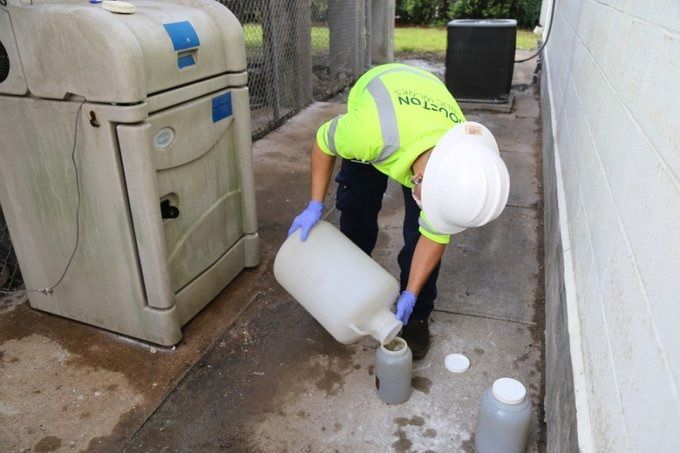
How much does virome prep influence our view of the human gut virome?
Short answer: a lot.
Long answer:
doi.org/10.1101/2025...
Different methods lead to distinct community structures, richness, and major virus-host abundance patterns.
🧵 1/5

How much does virome prep influence our view of the human gut virome?
Short answer: a lot.
Long answer:
doi.org/10.1101/2025...
Different methods lead to distinct community structures, richness, and major virus-host abundance patterns.
🧵 1/5
Meet MetaEdit: a platform for pathway-scale metagenomic editing inside the gut microbiome. science.org/doi/10.1126/...

Meet MetaEdit: a platform for pathway-scale metagenomic editing inside the gut microbiome. science.org/doi/10.1126/...
www.science.org/doi/10.1126/...

www.science.org/doi/10.1126/...
We identified a plasmid vector for the strain and generated a large RB-TnSeq library, screening for genes impacting plastic degradation.
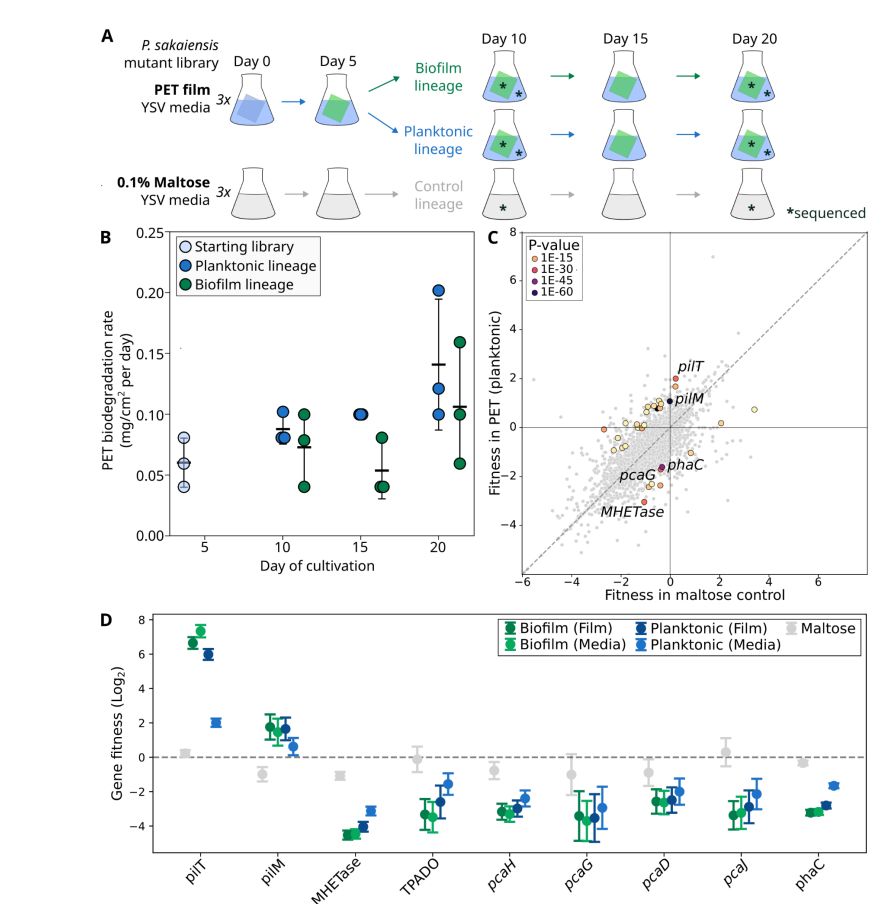
We identified a plasmid vector for the strain and generated a large RB-TnSeq library, screening for genes impacting plastic degradation.
vConTACT3 delivers a unified, scalable, and transparent framework for genome-based virus taxonomy — helping translate big viral data into systematic classification.
🔗 Read the preprint: doi.org/10.1101/2025...
Improvements details below 👇

vConTACT3 delivers a unified, scalable, and transparent framework for genome-based virus taxonomy — helping translate big viral data into systematic classification.
🔗 Read the preprint: doi.org/10.1101/2025...
Improvements details below 👇
We present a foundational genomic resource of human gut microbiome viruses. It delivers high-quality, deeply curated data spanning taxonomy, predicted hosts, structures, and functions, providing a reference for gut virome research. (1/8)
www.biorxiv.org/content/10.1...

We present a foundational genomic resource of human gut microbiome viruses. It delivers high-quality, deeply curated data spanning taxonomy, predicted hosts, structures, and functions, providing a reference for gut virome research. (1/8)
www.biorxiv.org/content/10.1...
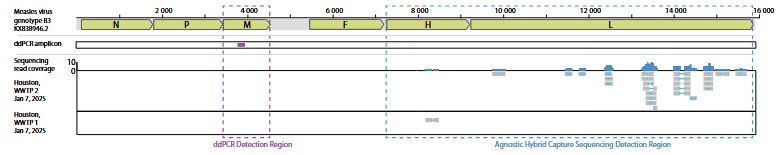
That is the question we asked in our latest pre-print.
Turns out you can.
1/
www.medrxiv.org/content/10.1...

That is the question we asked in our latest pre-print.
Turns out you can.
1/
www.medrxiv.org/content/10.1...
@natmicrobiol.nature.com today!
Fecal microbiota transplant (FMT), donor 💩 => patients' gut, is an effective treatment for recurrent C. difficile infection & is being evaluated for Inflammatory Bowel Diseases (IBD) & other conditions 1/
📄 rdcu.be/eL8mR

@natmicrobiol.nature.com today!
Fecal microbiota transplant (FMT), donor 💩 => patients' gut, is an effective treatment for recurrent C. difficile infection & is being evaluated for Inflammatory Bowel Diseases (IBD) & other conditions 1/
📄 rdcu.be/eL8mR
We sampled indoor air for a year in a Belgian daycare and used shotgun metagenomics to track viruses. We recovered many viral genomes of interest.
What would you sample next?
Read at @eurosurveillance.org
@emmanuel-microb.bsky.social, @jellematthijnssens.bsky.social
-->

We sampled indoor air for a year in a Belgian daycare and used shotgun metagenomics to track viruses. We recovered many viral genomes of interest.
What would you sample next?
Read at @eurosurveillance.org
@emmanuel-microb.bsky.social, @jellematthijnssens.bsky.social
-->
Massive thanks to Klara for driving this

Massive thanks to Klara for driving this
Nanopore's getting accurate, but
1. Can this lead to better metagenome assemblies?
2. How, algorithmically, to leverage them?
with co-author Max Marin @mgmarin.bsky.social, supervised by Heng Li @lh3lh3.bsky.social
1 / N
Nanopore's getting accurate, but
1. Can this lead to better metagenome assemblies?
2. How, algorithmically, to leverage them?
with co-author Max Marin @mgmarin.bsky.social, supervised by Heng Li @lh3lh3.bsky.social
1 / N
blast.ncbi.nlm.nih.gov/doc/blast-ne...
blast.ncbi.nlm.nih.gov/doc/blast-ne...

(both seem great)
#metaresearch #openscience 🧪
Learn more: plos.io/45wPZB9

(both seem great)
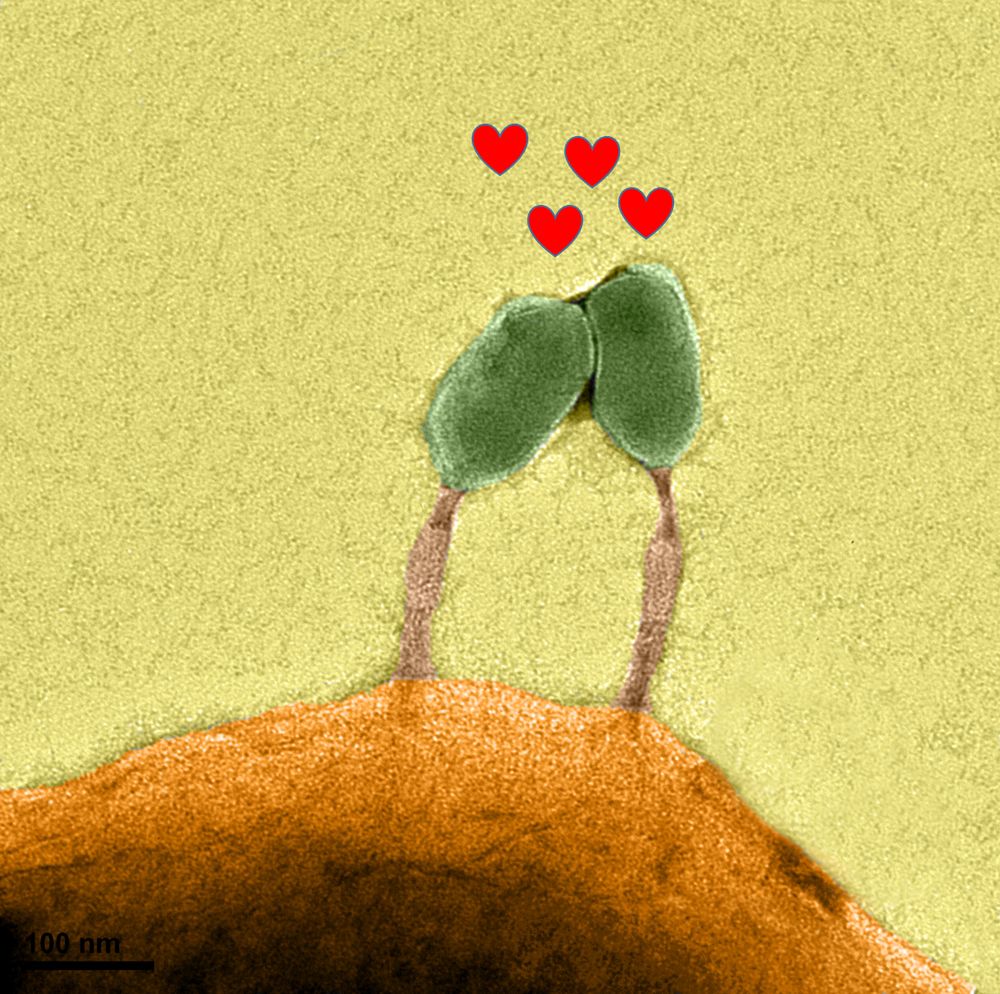

Led by @sherrynyeo.bsky.social, @erinmayc.bsky.social, and friends, we continue our journey to find viral DNA in our favorite place-- the overlooked and discarded reads in existing data! 1/

Led by @sherrynyeo.bsky.social, @erinmayc.bsky.social, and friends, we continue our journey to find viral DNA in our favorite place-- the overlooked and discarded reads in existing data! 1/
Apply by 2 July, details below!
💷 £36,720 to £39,750
🗓️ Apply by 2 July 2025
➡️ buff.ly/7M2mDaa

Apply by 2 July, details below!
The GlobDB is a database of "species dereplicated" microbial genomes, and as of release 226 contains twice the number of species-representative genomes (306,260) than the latest GTDB release.
The GlobDB is a database of "species dereplicated" microbial genomes, and as of release 226 contains twice the number of species-representative genomes (306,260) than the latest GTDB release.
myloasm-docs.github.io
myloasm-docs.github.io